PythonScript for protein colorizing
-
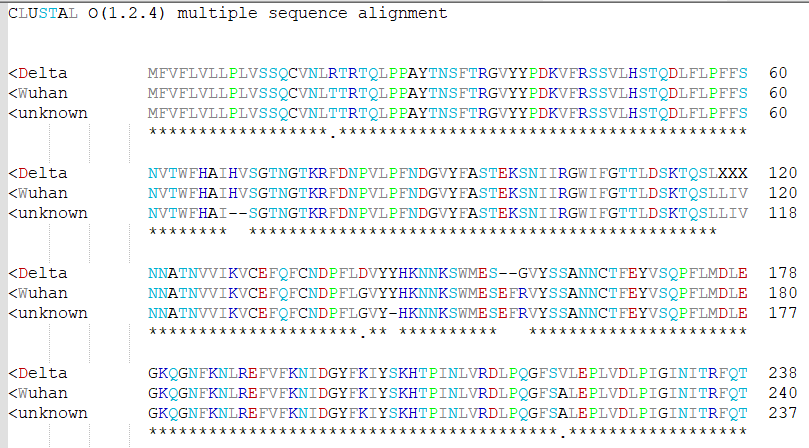
I thought it would be fun to make a lexer for colorizing protein sequences according to amino acid properties, and the current UDL system doesn’t seem well-suited for this. So I used PythonScript!
See https://github.com/molsonkiko/NPP_protein_lexer and maybe submit a pull request if you have an idea of how to improve it.
-
@Mark-Olson said in PythonScript for protein colorizing:
colorizing protein sequences according to amino acid properties,
I have an idea what you’re talking about but no experience in that field whatsoever. But between this and your JSONTools plugin, I’m happy to see a new talented developer contributing to the Notepad++ ecosystem.
Keep up the good work!
Cheers.
-
Just asking, I didn’t look at it: Does it absolutely need its own custom script, or could the EnhanceAnyLexer plugin have been used to do the same thing? This is typically the goto plugin when the UDL system falls short.
-
@Alan-Kilborn
I wasn’t really familiar with EnhanceAnyLexer prior to making this script, but now that I’ve tried it out this is actually really simple!Just need to create a lexer called
proteinwith white text background, install the plugin and add this toEnhanceAnyLexerConfig.ini[protein] ; default (black) 0x000000 = [^DEPHKRNQSTILMFWVACGY] ; acidic (blue) 0xbe0000 = [DE] ; cyclic (green) 0x00f600 = P ; basic (red) 0x0000be = [HKR] ; hydrophilic (cyan) 0xb0d000 = [NQST] ; amphiphilic (orange) 0x00b0d0 = [ACGY] ; lipophilic 0x848484 = [ILMFWV] ; check in the respective styler xml if the following IDs are valid excluded_styles = 1,2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,20,21,22,23and you get the same result, except better because amphiphilic amino acids are now orange.
-
@Mark-Olson And now I have to look up “amphophilic”
Neat
Hello! It looks like you're interested in this conversation, but you don't have an account yet.
Getting fed up of having to scroll through the same posts each visit? When you register for an account, you'll always come back to exactly where you were before, and choose to be notified of new replies (either via email, or push notification). You'll also be able to save bookmarks and upvote posts to show your appreciation to other community members.
With your input, this post could be even better 💗
Register Login